Self-Distillation Improves DNA Sequence Inference
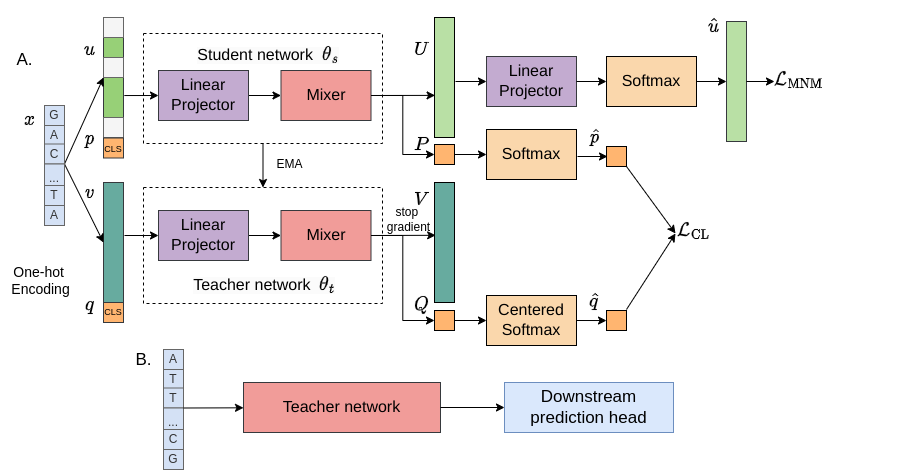
Self-supervised pretraining (SSP) has been recognized as a method to enhance prediction accuracy in various downstream tasks. However, its efficacy for DNA sequences remains somewhat constrained. This limitation stems primarily from the fact that most existing SSP approaches in genomics focus on masked language modeling of individual sequences, neglecting the crucial aspect of encoding statistics across multiple sequences. To overcome this challenge, we introduce an innovative deep neural network model, which incorporates collaborative learning between a `student' and a `teacher' subnetwork. In this model, the student subnetwork employs masked learning on nucleotides and progressively adapts its parameters to the teacher subnetwork through an exponential moving average approach. Concurrently, both subnetworks engage in contrastive learning, deriving insights from two augmented representations of the input sequences. This self-distillation process enables our model to effectively assimilate both contextual information from individual sequences and distributional data across the sequence population. We validated our approach with preliminary pretraining using the human reference genome, followed by applying it to 20 downstream inference tasks. The empirical results from these experiments demonstrate that our novel method significantly boosts inference performance across the majority of these tasks. Our code is available at https://github.com/wiedersehne/FinDNA.
PDF Abstract