Predicting Cellular Responses with Variational Causal Inference and Refined Relational Information
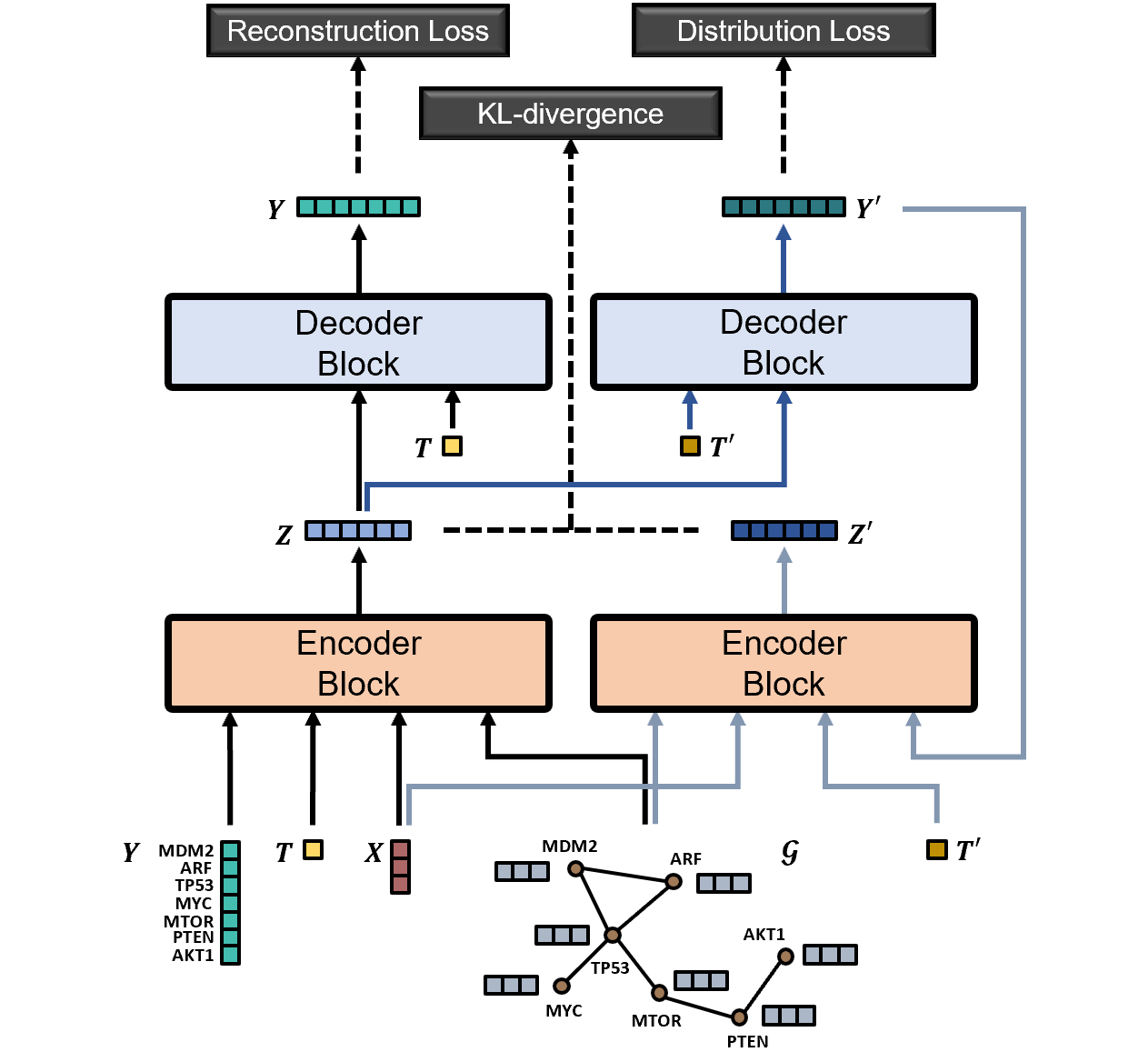
Predicting the responses of a cell under perturbations may bring important benefits to drug discovery and personalized therapeutics. In this work, we propose a novel graph variational Bayesian causal inference framework to predict a cell's gene expressions under counterfactual perturbations (perturbations that this cell did not factually receive), leveraging information representing biological knowledge in the form of gene regulatory networks (GRNs) to aid individualized cellular response predictions. Aiming at a data-adaptive GRN, we also developed an adjacency matrix updating technique for graph convolutional networks and used it to refine GRNs during pre-training, which generated more insights on gene relations and enhanced model performance. Additionally, we propose a robust estimator within our framework for the asymptotically efficient estimation of marginal perturbation effect, which is yet to be carried out in previous works. With extensive experiments, we exhibited the advantage of our approach over state-of-the-art deep learning models for individual response prediction.
PDF Abstract