Integrative Imaging Informatics for Cancer Research: Workflow Automation for Neuro-oncology (I3CR-WANO)
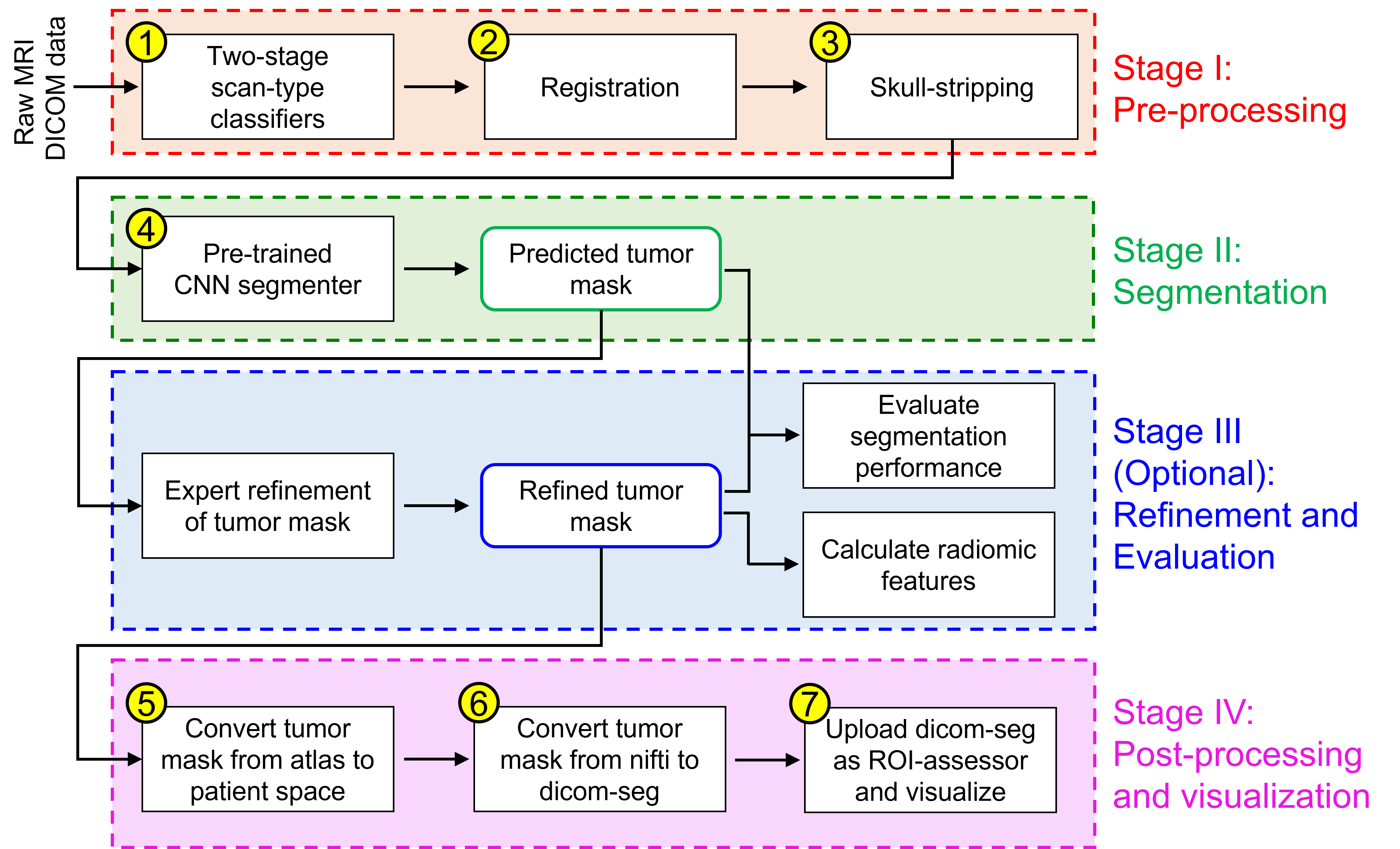
Efforts to utilize growing volumes of clinical imaging data to generate tumor evaluations continue to require significant manual data wrangling owing to the data heterogeneity. Here, we propose an artificial intelligence-based solution for the aggregation and processing of multisequence neuro-oncology MRI data to extract quantitative tumor measurements. Our end-to-end framework i) classifies MRI sequences using an ensemble classifier, ii) preprocesses the data in a reproducible manner, iii) delineates tumor tissue subtypes using convolutional neural networks, and iv) extracts diverse radiomic features. Moreover, it is robust to missing sequences and adopts an expert-in-the-loop approach, where the segmentation results may be manually refined by radiologists. Following the implementation of the framework in Docker containers, it was applied to two retrospective glioma datasets collected from the Washington University School of Medicine (WUSM; n = 384) and the M.D. Anderson Cancer Center (MDA; n = 30) comprising preoperative MRI scans from patients with pathologically confirmed gliomas. The scan-type classifier yielded an accuracy of over 99%, correctly identifying sequences from 380/384 and 30/30 sessions from the WUSM and MDA datasets, respectively. Segmentation performance was quantified using the Dice Similarity Coefficient between the predicted and expert-refined tumor masks. Mean Dice scores were 0.882 ($\pm$0.244) and 0.977 ($\pm$0.04) for whole tumor segmentation for WUSM and MDA, respectively. This streamlined framework automatically curated, processed, and segmented raw MRI data of patients with varying grades of gliomas, enabling the curation of large-scale neuro-oncology datasets and demonstrating a high potential for integration as an assistive tool in clinical practice.
PDF Abstract